Up to this point, you have been keeping your Python code and Markdown comments in a single Jupyter notebook document. This is great for exploration and learning. However, there are cases where you want to have Python code in a separate document. When you have long Python code blocks or a set of functions used across many different notebooks, moving them out makes your notebook much easier to read and use.
An alternative to typing all your commands directly into cells is to list them in a Python script file. A Python script file is simply a plain text file containing a list of the commands you want to run, formatted exactly as if you were to type them into a cell. Python script files traditionally use the .py file extension.
1. Creating a script file¶
Because a Python script file is simply a list of commands, we can easily create a basic one and test things out right inside JupyterLab.
First, create a new text file by clicking on File > New > Python File in the JupyterLab menu bar. You can do the same in VS Code (File > New File > Python File) or Colab (File > Download > Download .py).
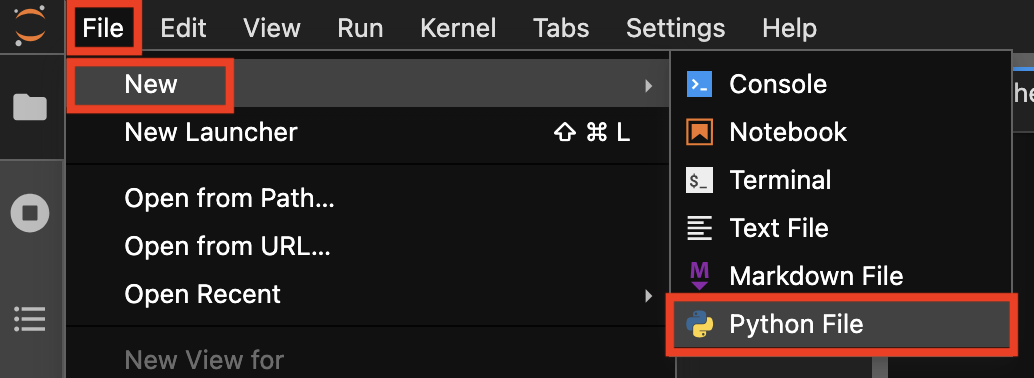
Creating a new Python file in JupyterLab.
This creates a new tab in your window with a blank slate. By default, new Python files will have the name untitled.py. You can rename the file by right clicking on it in the file browser and renaming it to spatial_tools.py.
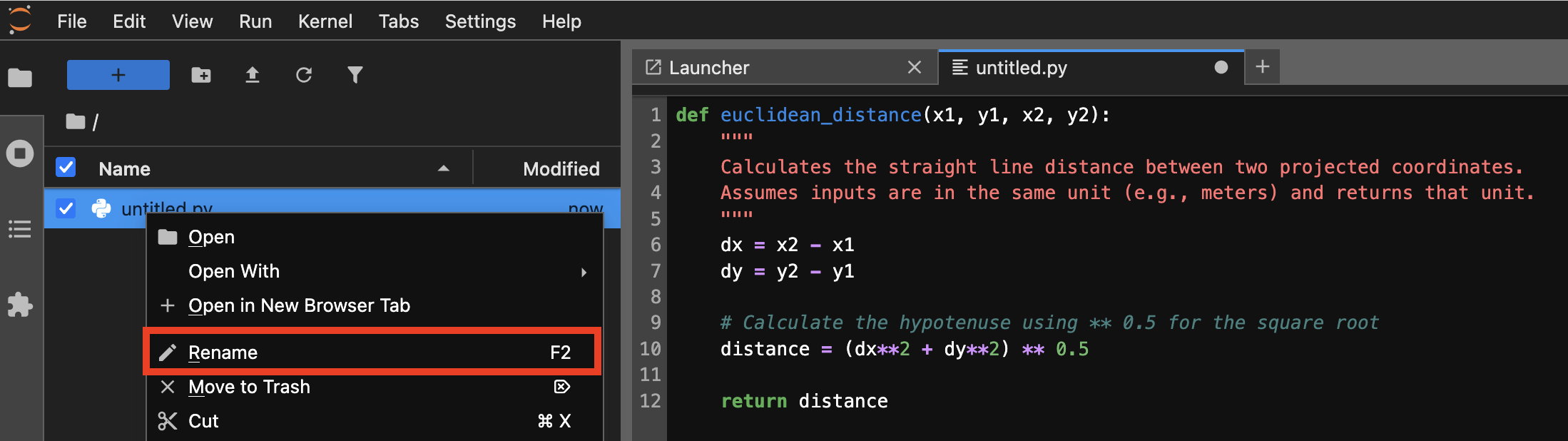
Renaming the new file in JupyterLab.
Start by copying and pasting the text below into your new editor panel. This is a function that calculates the straight line Euclidean distance between two projected coordinates using the basic math operators you learned in Lesson 2:
def euclidean_distance(x1, y1, x2, y2):
"""
Calculates the straight line distance between two projected coordinates.
Assumes inputs are in the same unit (e.g., meters) and returns that unit.
"""
dx = x2 - x1
dy = y2 - y1
# Calculate the hypotenuse using ** 0.5 for the square root
distance = (dx**2 + dy**2) ** 0.5
return distance
Be sure to save your spatial_tools.py file after making your changes. You have just created your first custom Python module!
2. The working directory¶
To use the function you just saved, your new Jupyter notebook needs to know where to find the script file. Python looks for imported files in your current working directory.
Hopefully you saved your spatial_tools.py file in the exact same folder as the new notebook you are currently working in. We can verify this running an IPython magic command called %ls in a code cell. The % symbol is a special built-in shortcut used to perform helpful tasks that are outside of standard Python code.
%ls
The %ls command lists all the files located in the directory where you are currently working. If you see spatial_tools.py in the output list, you are all set to proceed.
Concept check¶
Imagine your file browser looks like this:
project_folder/
├── analysis.ipynb <-- You are working here
└── scripts/
└── my_code.py
If you run %ls in analysis.ipynb, will you see my_code.py? Will a standard import my_code work?
Check your understanding
No and No.
%lslists files in the current directory (project_folder). It will show thescriptsfolder, but not the contents inside it.Therefore, a standard
import my_codewill fail because Python cannot see the file sitting in the subfolder. To fix this, you would need to movemy_code.pyup intoproject_folder.
3. Importing script functions¶
Now that your notebook and your script are in the same folder, you can access your custom functions using an import statement.
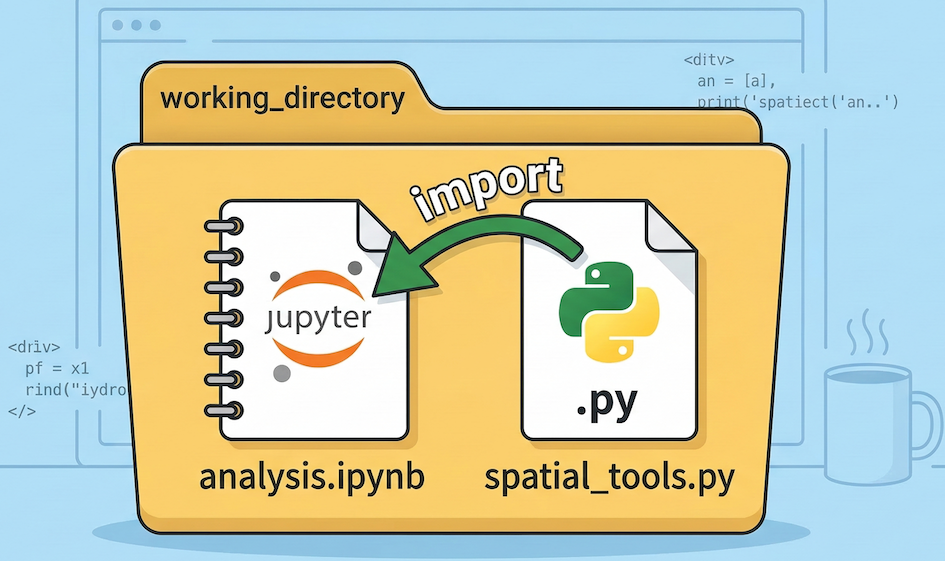
Python looks for the imported script file in the same working directory as your active notebook.
Let us import our euclidean_distance() function from the script file:
from spatial_tools import euclidean_distance
Now we can use the function to see that it is working. Let us calculate the distance between Zurich and Bern using their Swiss Grid (LV95) coordinates in meters:
# Zurich: 2683000, 1248000 | Bern: 2600000, 1200000
dist_meters = euclidean_distance(2683000, 1248000, 2600000, 1200000)
dist_km = dist_meters / 1000
print(f"The straight line distance is {dist_km:.1f} km")
Other ways to import¶
It is often useful to import the whole script and all of its functions at once. This can be done by importing the file and giving it a short alias.
import spatial_tools as st
dist_meters = st.euclidean_distance(2683000, 1248000, 2600000, 1200000)
Concept check¶
Look at these two different ways to import the same function. Predict how you would call the function in the cell immediately following the import.
Scenario A:
from spatial_tools import euclidean_distance
# How do I call it here?
Scenario B:
import spatial_tools as st
# How do I call it here?
Scenario A or B?
Scenario A: You brought the function directly into your notebook’s namespace. You call it simply by its name:
euclidean_distance(...)
Scenario B: You imported the whole module as an object named st. You must use “dot notation” to access the function inside that object:
st.euclidean_distance(...)
4. Building a modular tool¶
Let us add a slightly more complex tool to our script. We will create a wrapper function that accepts coordinate pairs grouped in lists, rather than forcing the user to type out four separate numbers.
Go back to your spatial_tools.py tab, leave two blank lines below your first function, and add the following code:
def calculate_distance(pt1, pt2, method="euclidean"):
"""
Wrapper function to calculate distance between two points.
Parameters
----------
pt1: <list>
A list containing [x, y] coordinates
pt2: <list>
A list containing [x, y] coordinates
method: <str>
The calculation method. Supported values: 'euclidean'
Returns
-------
<float>
Computed distance.
"""
if method == "euclidean":
# Extract coordinates from the lists and pass to our dedicated function
dist = euclidean_distance(pt1[0], pt1[1], pt2[0], pt2[1])
else:
print("Method not supported yet.")
dist = None
return dist
Save the file once more. Your custom module now contains two functions.
5. The notebook kernel trap¶
Now that we have updated our script, we want to test the new calculate_distance function in our notebook. However, when working in a Jupyter Notebook, reloading updated modules can be tricky.
To save memory, Python only imports a script once per session. If you change a script file and save it, simply running the import cell again will not load your new changes. Python thinks it already has the file and ignores the command.
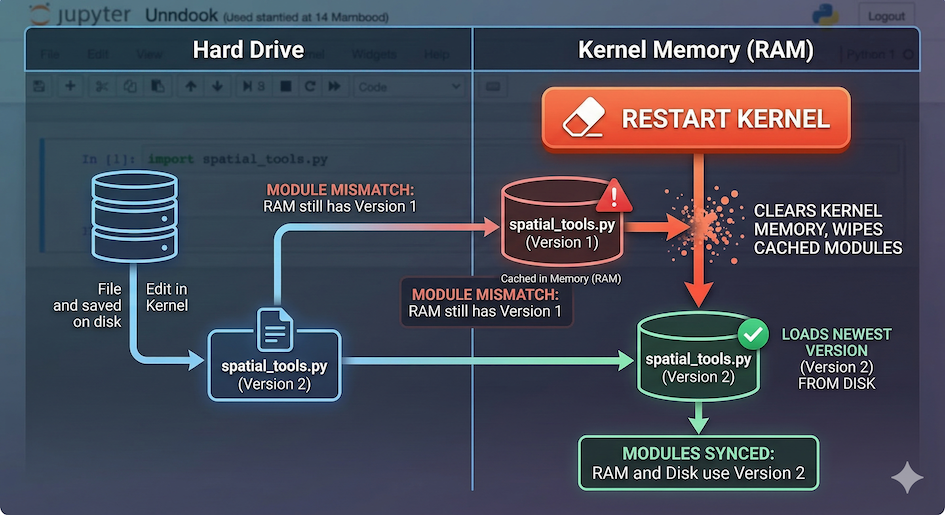
When a script is first imported, it is loaded into the Kernel’s active memory. Updating the text file on your hard drive does not update the memory until the Kernel is explicitly restarted.
The easiest way to force Python to see your updates is to restart the IPython kernel.
Go to Kernel > Restart kernel... in the menu bar.
Wait a moment for the memory to clear.
Re-run your import cell.
After restarting the kernel, you can safely import your updated script and use your new tool:
import spatial_tools as st
zurich = [2683000, 1248000]
bern = [2600000, 1200000]
distance_m = st.calculate_distance(zurich, bern, method="euclidean")
print(f"Distance: {distance_m / 1000:.1f} km")
6. Exercises¶
To practice working with script files, complete the following tasks using your spatial_tools.py script.
Focus on understanding the workflow between your script and your notebook.
Exercise 1: Add a Manhattan distance function¶
Open your spatial_tools.py file in the text editor.
Write a new function called
manhattan_distance(x1, y1, x2, y2)that calculates the grid-based distance between two coordinates.Remember from Lesson 2 that you can get the absolute (positive) difference by squaring it and taking the square root:
((x2 - x1)**2)**0.5.Save the file.
# Write your code here
Sample solution (spatial_tools.py)
def manhattan_distance(x1, y1, x2, y2):
"""Calculates the grid-based distance between two projected coordinates."""
dx = ((x2 - x1)**2)**0.5
dy = ((y2 - y1)**2)**0.5
distance = dx + dy
return distanceKey idea: You are defining a new, isolated tool in your script. It takes the exact same four inputs as the Euclidean function, but applies a different mathematical logic.
Exercise 2: Update the wrapper function¶
Stay in your spatial_tools.py file.
Update your existing
calculate_distancefunction to support the new method.Add an
elif method == "manhattan":block that extracts the coordinates from thept1andpt2lists and passes them to your newmanhattan_distancefunction.Save the file again.
# Write your code here
Sample solution (spatial_tools.py)
def calculate_distance(pt1, pt2, method="euclidean"):
"""Wrapper function to calculate distance between two points."""
if method == "euclidean":
dist = euclidean_distance(pt1[0], pt1[1], pt2[0], pt2[1])
elif method == "manhattan":
dist = manhattan_distance(pt1[0], pt1[1], pt2[0], pt2[1])
else:
print("Method not supported yet.")
dist = None
return distKey idea:
The wrapper function acts as a single, user-friendly interface. It routes the user’s request to the correct underlying mathematical function based on the method argument.
Exercise 3: Test your updates¶
Return to your Jupyter notebook.
Restart the kernel to clear the memory.
Import your updated module.
Test both the Euclidean and Manhattan functionality to see how much longer the grid-based path is compared to the straight-line path.
# Write your code here
Sample solution (Jupyter Notebook)
import spatial_tools as st
zurich = [2683000, 1248000]
bern = [2600000, 1200000]
euclid_dist = st.calculate_distance(zurich, bern, method="euclidean")
manhat_dist = st.calculate_distance(zurich, bern, method="manhattan")
print(f"Euclidean path: {euclid_dist / 1000:.1f} km")
print(f"Manhattan path: {manhat_dist / 1000:.1f} km")Key idea:
Restarting the kernel is crucial. It ensures Python loads the newest saved version of your .py script into memory instead of reusing the old one.
7. Summary¶
In this section, you learned how to move your code out of the notebook sandbox and into reusable script files. You discovered how to:
Create and save a
.pyfile containing custom functions.Verify your working directory using
%ls.Import specific functions or entire scripts into your notebook.
Safely reload updated scripts by restarting the IPython kernel.
By writing modular tools in separate scripts, you keep your main notebooks clean, readable, and focused entirely on data analysis.
What comes next?¶
You have now built and imported your own custom module. But you do not have to write every tool from scratch! In the next section, we will explore the Python Standard Library and discover how to use the exact same import syntax to unlock powerful tools that are already built into Python.